 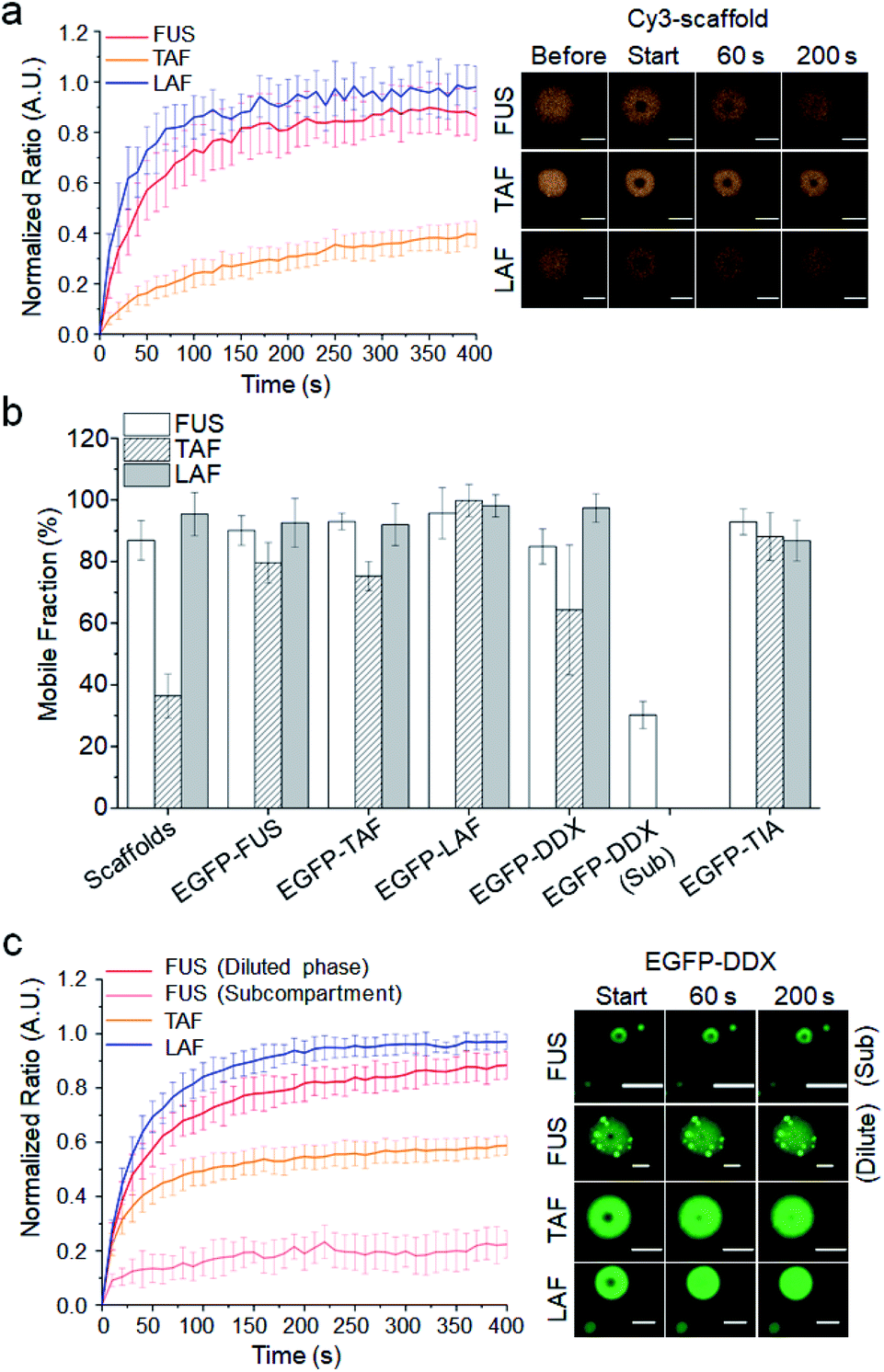
Aberrant phase transition of TDP-43 and FUS leads to protein aggregation and disrupts their regular cell function. We also review evidences that connect the phase separation property of TDP-43 and FUS to their functional roles in cells. In this article, we review the factors that mediate and regulate phase separation of TDP-43 and FUS. They have similar domain structures that provide multivalent interactions driving their phase separation in vitro and in the cellular environment. TDP-43 and FUS are two such RNA-binding proteins that mislocalize and aggregate in patients of ALS and FTD. However, aberrant phase transition of these proteins leads to the formation of insoluble protein aggregates, which are pathological hallmarks of neurodegenerative diseases including ALS and FTD. Liquid-liquid phase separation of RNA-binding proteins mediates the formation of numerous membraneless organelles with essential cellular function. Department of Biochemistry and Molecular Biology, Thomas Jefferson University, Philadelphia, PA, United States.The length of the protein (amino acid residues according to Ensembl), molecular mass (kDalton), predicted signal peptide (according to a majority of the signal peptide predictors SPOCTOPUS, SignalP 4.0, and Phobius) and the number of predicted transmembrane region(s) (according to MDM) are also reported. The Gene Ontology terms assigned to this protein are listed if expanding the Gene ontology column. Parent protein classes are in bold font and subclasses are listed under the parent class. The protein classes assigned to this protein are shown if expanding the data in the protein class column. The data in the UniProt column can be expanded to show links to all matching UniProt identifiers for this protein. The ENSP identifier links to the Ensembl website protein summary, while the ENST identifier links to the Ensembl website transcript summary for the selected splice variant. The protein information section displays alternative protein-coding transcripts (splice variants) encoded by this gene according to the Ensembl database. Common (purple) and unique (grey) regions between different splice variants of the gene are also displayed ( read more), and at the bottom of the protein view is the protein scale. 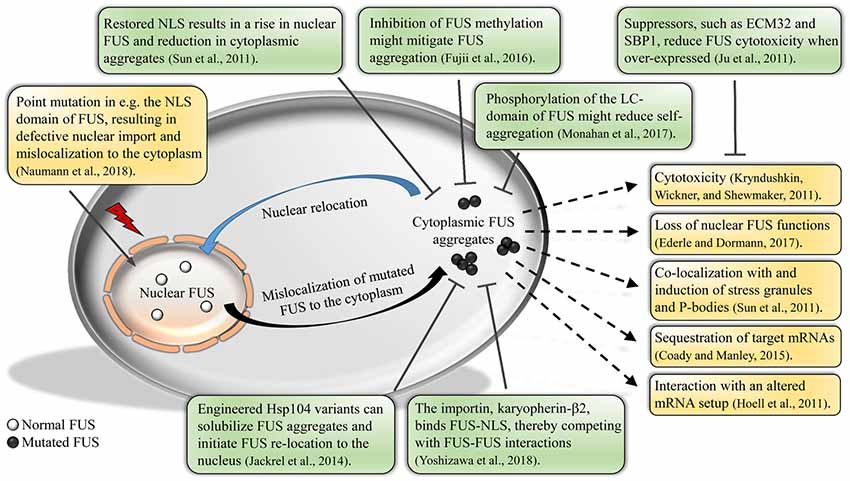
Low complexity regions are shown in yellow and InterPro regions in green. If a signal peptide is predicted by a majority of the signal peptide predictors SPOCTOPUS, SignalP 4.0, and Phobius (turquoise) and/or transmembrane regions (orange) are predicted by MDM, these are displayed. the tendency for different regions of the protein to generate an immune response, with peak regions being predicted to be more antigenic.The curve shows average values based on a sliding window approach using an in-house propensity scale. The curve in blue displays the predicted antigenicity i.e. The region with the lowest possible identity is always selected for antigen design, with a maximum identity of 60% allowed for designing a single-target antigen ( read more). A yellow triangle on the bar indicates a <100% sequence identity to the protein target.īelow the antigens, the maximum percent sequence identity of the protein to all other proteins from other human genes is displayed, using a sliding window of 10 aa residues (HsID 10) or 50 aa residues (HsID 50). 
The tabs at the top of the protein view section can be used to switch between the different splice variants to which an antigen has been mapped.Īt the top of the view, the position of the antigen (identified by the corresponding HPA identifier) is shown as a green bar. The protein browser displays the antigen location on the target protein(s) and the features of the target protein. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
